Introduction: A hands-on kitchen moment with a scanner
I once watched a colleague treat a new imaging rig like a mystery stew—she poked, prodded, and sighed until the colors finally came alive on screen. The room smelled faintly of metal and warm plastic; the display glowed like a simmering pot. In vivo imaging shows that tiny living systems can tell big stories, but only when the recipe is right. Data from routine lab checks — throughput dips, signal drift, phantom mismatches — stack up faster than you’d expect. So how do we stop burning the dish and start serving consistent, useful images?
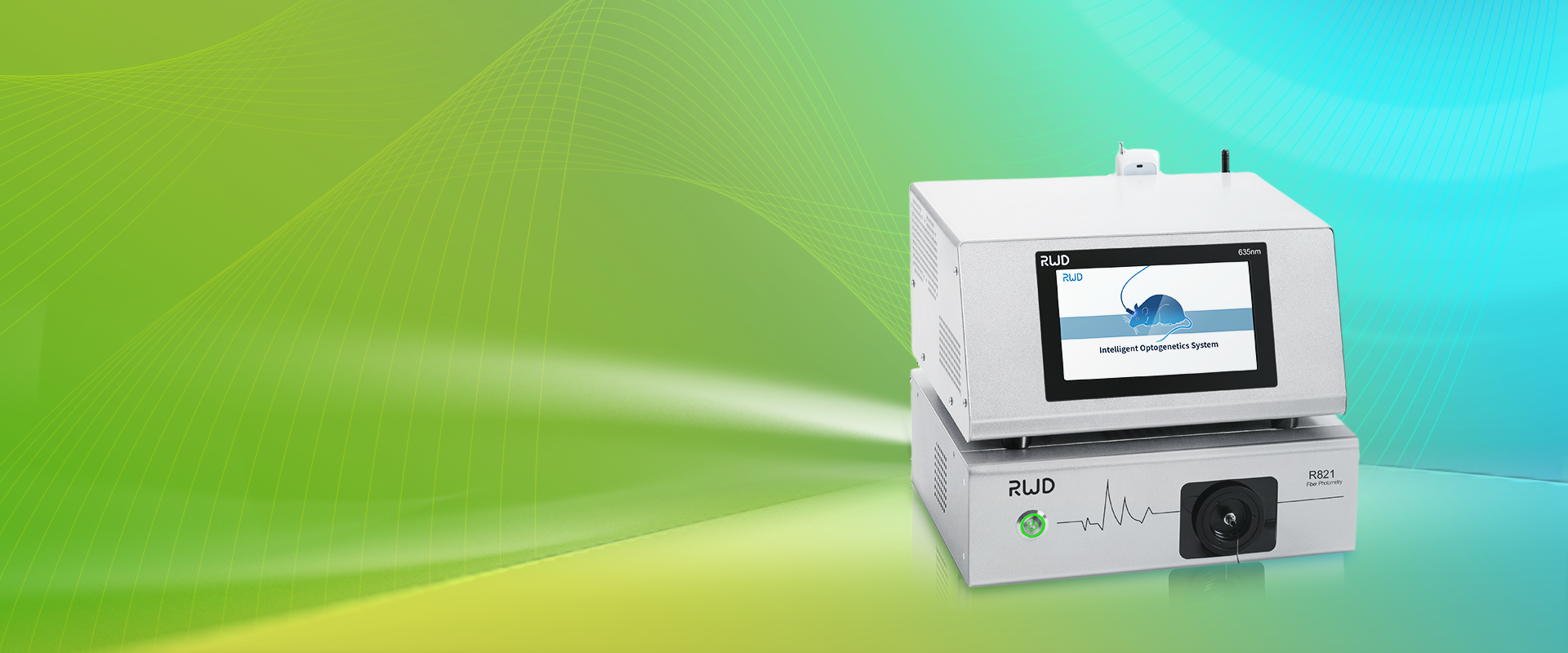

I write this as someone who has scrubbed scanners at dawn and debugged bad contrasts at midnight. I like to think of systems the way a chef thinks about knives: the right edge, kept sharp, makes work joyful. You’ll see terms like calibration phantom and photon counting pop up below — practical tools, not jargon. (Also — I waste fewer reagents when I follow a checklist.) Let’s move from that kitchen to the bench and look at what really trips teams up next.
Part 2 — Deeper layer: Why traditional fixes often fail
What goes wrong?
I’ll be blunt: many labs buy a small animal in vivo imaging system and expect it to be turnkey. Reality bites. First, global calibration routines hide local errors. A single calibration phantom run cannot catch temperature gradients or detector aging. Second, data pipelines assume neat, labeled input. They don’t handle messy metadata, and so crucial frames get dropped. Third, hardware and software mismatches — firmware that needs power converters tuned or algorithms that assume a photon counting detector — create subtle biases that show up days later. Look, it’s simpler than you think to miss these details, yet the impact compounds in measurements and publications.
From my hands-on work, I’ve seen two repeat patterns. One: labs conflate spatial resolution losses with contrast issues, then chase the wrong fix. Two: teams patch software with quick filters that hide noise but also erase weak signals — that’s disastrous for longitudinal studies. Fluorescence tomography can be forgiving, but only if the acquisition, calibration phantom checks, and reconstruction settings align. If they don’t, you get pretty pictures that are scientifically weak. I prefer breaking the problem into small tests: one variable at a time, repeatable, logged. That discipline saves weeks.
Part 3 — Forward-looking: Principles for better systems and decisions
What’s Next: Practical principles, not buzzwords
New technology should solve real pain, not create bookkeeping work. For the next generation of small animal in vivo imaging system designs, I focus on modularity and transparency. Modular detectors mean you can swap a photon counting module without revalidating the whole pipeline. Transparent metadata — machine-readable logs about temperature, gain, and firmware — lets you spot drift early. Edge computing nodes that preprocess raw frames at acquisition reduce storage load and speed QC checks. These aren’t sci‑fi ideas; they’re practical shifts that cut troubleshooting time in half, if implemented thoughtfully.
Another principle: design for reproducibility. That means built-in calibration phantom routines, versioned reconstruction code, and clear user workflows. Don’t expect users to become programmers overnight — provide sensible defaults and visible knobs. I’ve argued (sometimes loudly) for vendor-supplied, validated pipelines so smaller labs can produce reliable longitudinal data without a full software team. — funny how that works, right? Ultimately, the right mix of hardware clarity and software hygiene will free researchers to ask better biological questions instead of babysitting instruments.

To close with something practical: when you evaluate systems, focus on three metrics — calibration repeatability (how little the system drifts across weeks), metadata completeness (are temperature, gain, and firmware logged automatically?), and end-to-end reproducibility (can an independent group reproduce an image using your raw data and settings?). Those three lenses tell you more than flashy specs. I stand by these priorities because I’ve seen them cut wasted experiment days in half. For labs looking to take the next step, consider suppliers who make validation easy and transparent. For reliable solutions and support, check BPLabLine.