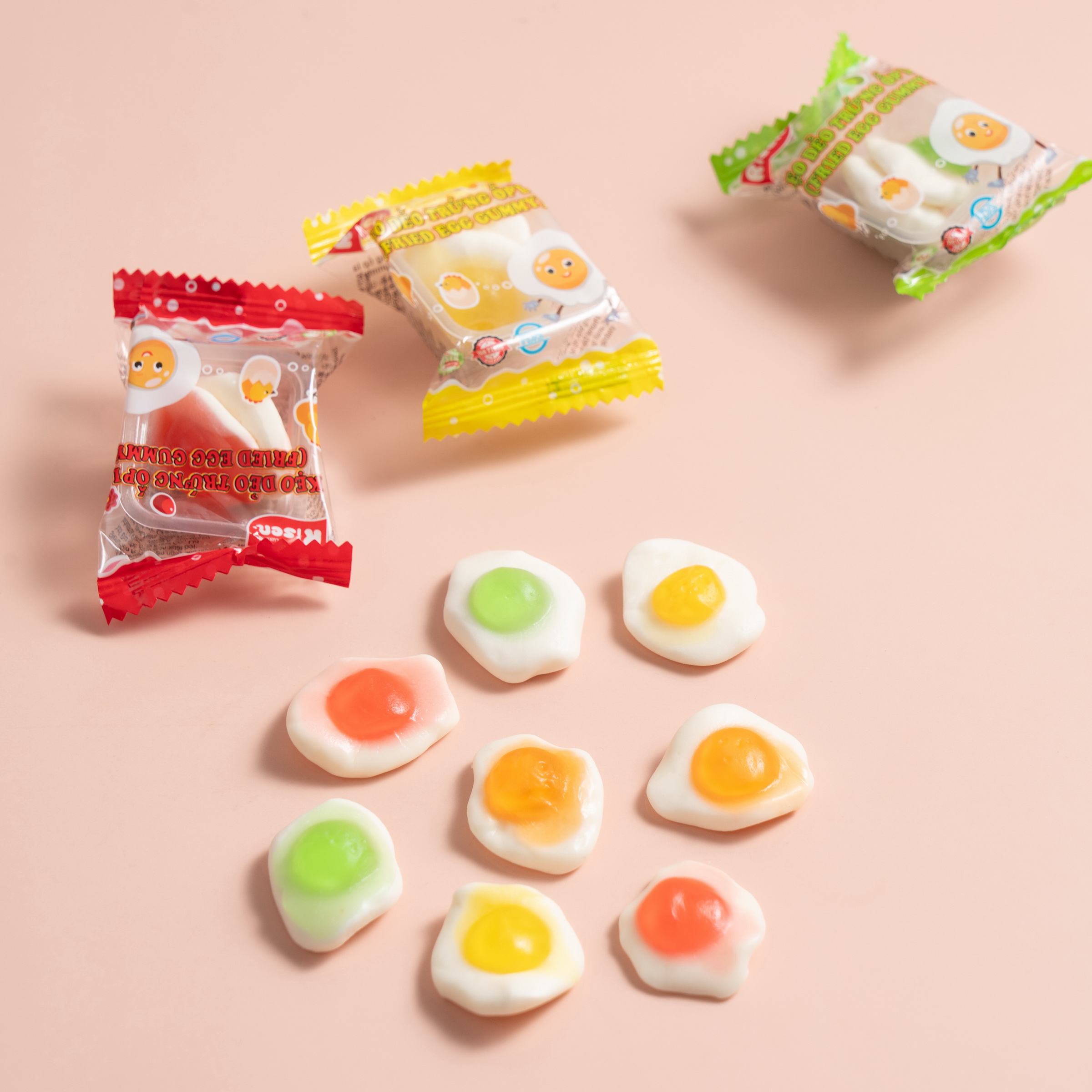
Introduction
I once sat in a lab at midnight watching a run finish — and I swore we’d do better next time. automated nucleic acid extraction is supposed to save time, but often it just swaps one bottleneck for another. Data shows many labs lose hours per week to manual cleanups and failed runs (yes, real numbers — like dozens of samples wasted). So what actually trips us up when we buy a machine and expect miracles? Let me walk you through what I see, what I’ve fixed, and what you can test next. — moving on to the deeper stuff.
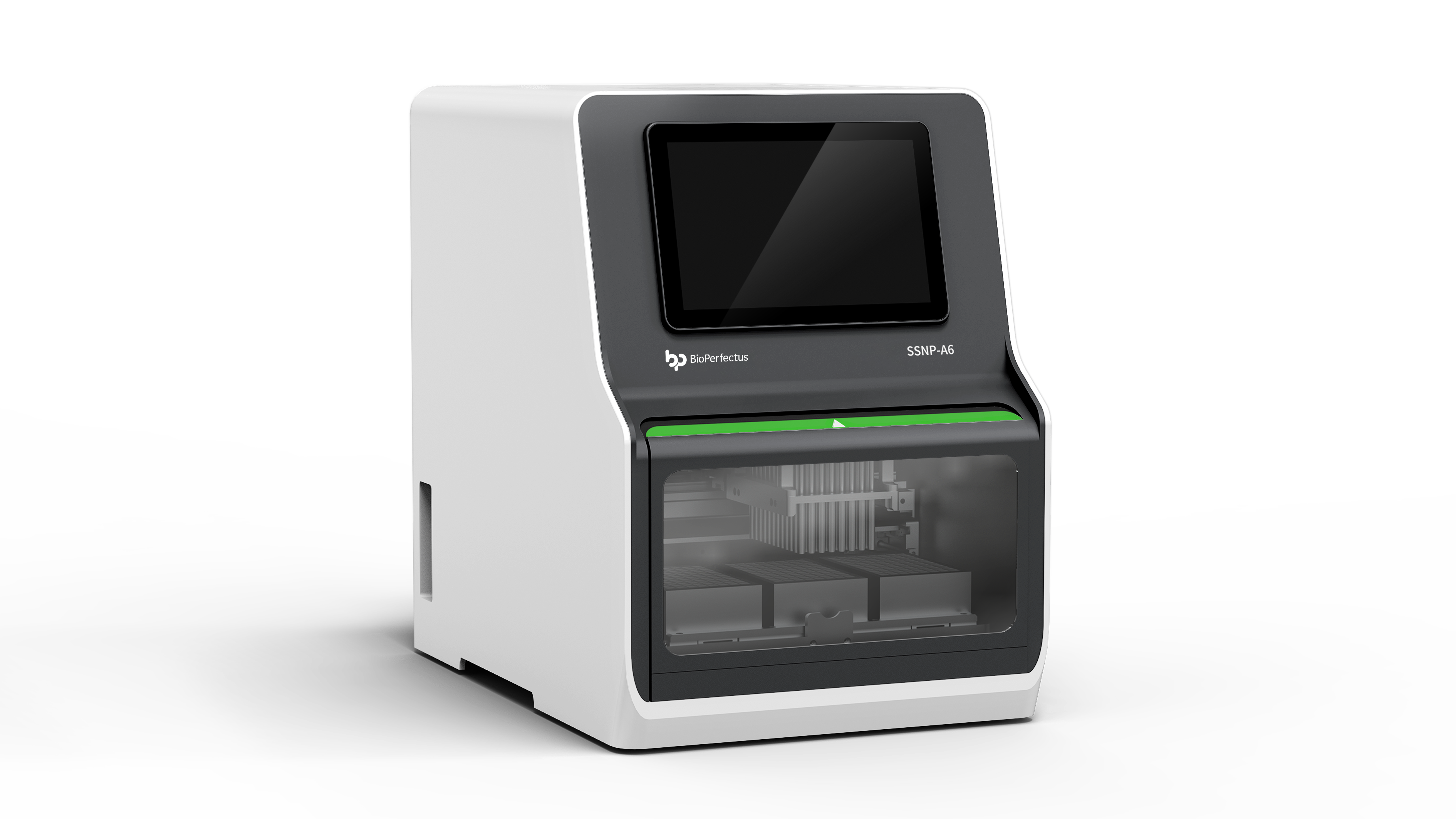

Where the usual solutions fall short
automated dna extraction machine often promises hands-off runs and perfect yields, but reality is messier. I’ve tested systems that choke on viscous samples, that lose RNA purity after a few batches, and that quietly reduce throughput because of long reagent change times. Magnetic bead separation steps can be sensitive to small timing shifts. Lysis buffer chemistry varies by sample. Throughput claims on data sheets rarely match a real schedule where setup, pauses, and QC checks matter. In short: the machine may be fast in a brochure, but your workflow isn’t. Look, it’s simpler than you think — check cycle layout and sample prep first.
Why does this happen?
Two main causes: hardware limitations and workflow mismatch. Pumps and valves age. Temperature control drifts. Software scripts assume ideal samples. We mismatch sample types and protocols (saliva vs. blood; low vs. high biomass). The result is variable RNA purity, repeated cleanups, and wasted kit reagents. I’ve seen labs lose confidence in automation because they didn’t tune protocols to their specimen types. You can fix many issues by adjusting bead ratios, changing wash volumes, or altering incubation times. Small tweaks. Big impact.
Looking ahead: principles and practical upgrades
I want to be practical here — not dreamy. New tech principles that matter: closed-loop feedback, modular reagent packs, and smart scheduling. If you pick systems that report real-time metrics (pressure, temperature, magnetic field strength), you get early warnings before a run fails. A modern automated dna extraction machine should let you swap modules fast and should support custom scripts for edge cases. That makes upgrades less painful and yields more consistent results.
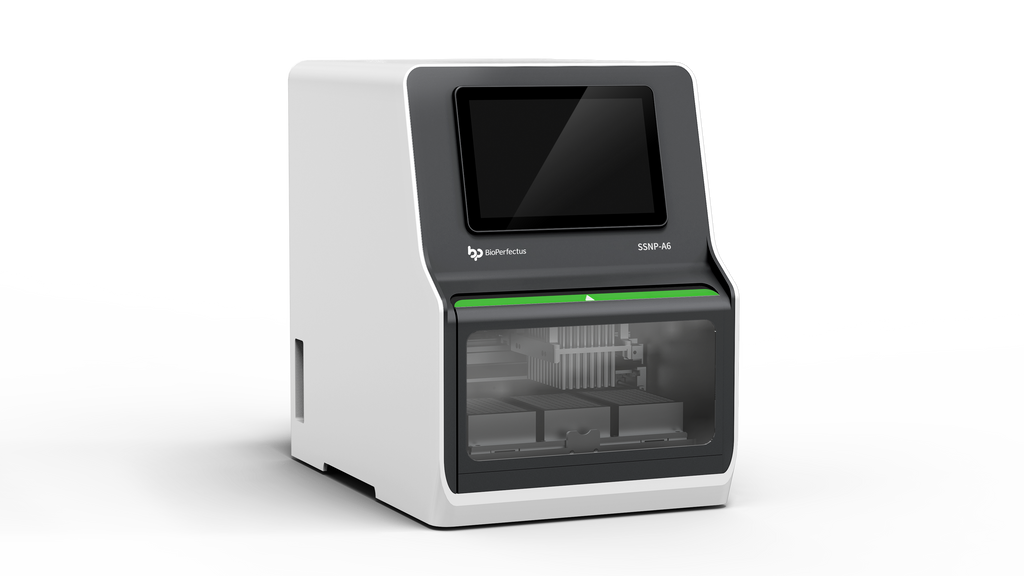

What’s Next?
Case example: we shifted one lab from a single monolithic unit to a modular line. Result — fewer aborted runs, higher RNA yield consistency, and a 25% cut in hands-on time. — funny how that works, right? Future outlook: expect more on-device QC and better reagent formats so labs spend less time troubleshooting and more time analyzing data.
How to choose — three practical metrics
When I advise teams, I tell them to test three things before buying: (1) Real throughput with your sample type — not the datasheet number; (2) Error transparency — can the system tell you why a sample failed?; (3) Protocol flexibility — can you edit critical steps like magnetic bead timings or wash volumes? These metrics predict day-to-day usability far better than glossy brochures. If a vendor can’t demo all three with your samples, don’t buy yet. We learned that the hard way — and saved time (and budget) after switching to more transparent systems.
Finally, if you want a reliable partner that provides clear specs and real demos, check solutions like BPLabLine. I’ve worked with teams who found the right fit there — and it changed their daily workflow for the better.